The beginnings of the The AI Dementia Imaging project
The project began when the Alan Turning Institute facilitated a meeting between Research Computing Services and Professor Zoe Kourzi – professor in Psychology at the University of Cambridge.
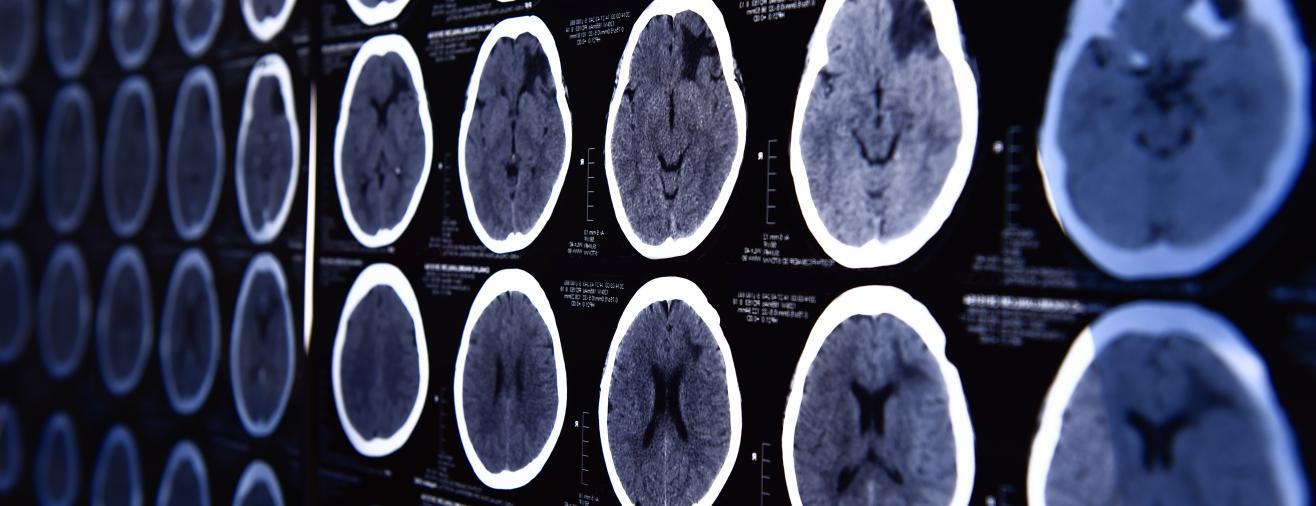
Professor Kourzi’s team was working on a machine learning algorithm that would deploy a proven research concept as a digital application that could be used in a clinical setting. Using brain scans from patients who went on to develop Alzheimer’s, their machine learning algorithm learnt to spot structural changes in the brain. When combined with the results from standard memory tests, the algorithm was able to provide a prognostic score – that is, the likelihood of the individual having Alzheimer’s disease. Currently, it can take several scans and tests to diagnose the disease. For those patients presenting with mild cognitive impairment – signs of memory loss or problems with language or visual/spatial perception – the algorithm was over 80% accurate in predicting those individuals who went on to develop Alzheimer’s disease. It was also able to predict how fast their cognition will decline over time.
The collaboration process
When Professor Kourzi and her team met with the Research Software Engineering team at Research Computing Services it was decided that the best way to deploy the algorithm in the clinic would be by using XNAT, and to package that application as a plug-in. XNAT was chosen because an instance of it is deployed in most NHS and research-imaging sites – it is an open source tool to allow clinicians to access, view and process imaging data for a cohort of patients. It is has a user interface that is easy to navigate for clinicians and that allows research ideas to be implemented in a secure way (crucial when dealing with sensitive data).
The Research Software Engineering team packaged software written by Professor Kourzi’s team in a plug-in for XNAT. It is possible to extend XNAT functionality with plug-ins – for instance, the fsPET plugin, which gives quantitative information about amyloid build-up in the brain from PET images. The plug-in proposed for this project was designed to do something similar. In addition to plugins, with XNAT it is also possible to create data types where user defined information (such as whether a patient has cognitive decline) is stored for the clinician to see and possibly to comment on.Designing the pipeline
To run the application, it was assumed that the project would have many patients with up to several scans (probably multiple MRI scans taken over a certain time period).The pipeline was designed to run in a three step process:Pull the cognitive metrics score associated with a patient from a REDCAP database
Retrieve an MRI image associated with the patient and call the ‘get probability map’ function
Given this scan’s probability map, the application would compute a grey matter score (GM score) which it would then combine with a patient’s cognitive scores (sourced from the REDCap database). This would produce an ‘ADscore’ – a value indicating any sort of cognitive impairment as some distance away from the ‘norm’. The result would then be appended to the cognitive score datatype
Facing challenges
The main engineering challenge for the RSE team was to enable clinicians to run a prediction pipeline by interacting with the XNAT user interface. The team packaged the steps above in a docker container (docker containers are the barebones of what you need to execute software on a system – including all necessary dependencies like matlab). The docker container had the matlab spm12 toolbox installed for accomplishing the MRI preprocessing, as well as scripts for pulling cognitive data from a remote database (REDCap). It was also designed with a python pipeline to take the results of step a. and b. above to give an ‘ADScore’.
The future of the project
Although the algorithm has been optimised to look for signs of Alzheimer’s disease, Professor Kourtzi and colleagues are now training it to recognise different forms of dementia, each of which has its own characteristic pattern of volume loss.
Dr Timothy Rittman from the Department of Clinical Neurosciences and a consultant at Addenbrooke’s Hospital, part of Cambridge University Hospitals (CUH) NHS Foundation Trust, is now leading a trial to look at whether this approach is useful in a clinical setting.
To date around 80 patients have taken part in the trial, which was run by CUH, Cambridgeshire and Peterborough NHS Foundation Trust and two NHS trusts in Brighton.
There are currently very few drugs available to help treat dementia. One of the reasons that clinical trials often fail is because it is thought that once a patient has developed symptoms, it may be too late to make a major difference. Having the ability to identify individuals at a very early stage could therefore help researchers develop new medicines.
If the trial is successful, the algorithm could be rolled out to thousands more patients across the country.
You can learn more about this project by vising the Alan Turing Institute site
.